Poster 1051: WES and RNAseq Comparison of PDX Mutations
Mutation Detection in Patient-Derived Xenografts by Whole Exome Sequencing and Transcriptome Sequencing
Wubin Qian, Jia Xue, Henry Q. Li, and Sheng Guo
Patient-derived xenografts (PDX) are widely used, highly predictive, preclinical oncology models, useful for a range of study types including drug efficacy evaluation, biomarker discovery and validation, drug mechanism and resistant mechanism studies, and tumor biology studies.
A good quality, well characterized PDX library should include genomic, pharmacology, and histology data, as well as clinical information associated with the original patient tumors. Various technologies are used for PDX genomic profiling e.g. RNAseq to obtain gene expression, RNA-level mutation, and gene fusion data, and whole exome sequencing (WES) for inferring DNA-level mutations and gene copy numbers.
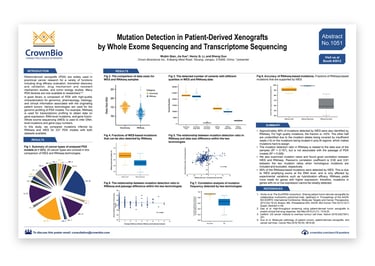
This poster compares PDX mutations inferred by RNAseq and WES for a panel of over 230 models, covering 20 cancer types.
Read this Poster to Discover:
- That approximately 48% of mutations detected by WES were also identified by RNAseq, with other mutations being unidentified due to the mutation alleles being covered by insufficient reads or the mutations being located in poly-N regions
- That 99% of RNAseq-based mutations were detected by WES, due to WES amplifying exons at the DNA level
- That mutation ratios show good correlation between WES and RNAseq
Download the Poster Now!



